|
soutenance d'HDR
Par Claude Loverdo
Le 5 Septembre 2022 à 14h00 - C404
|
Résumé
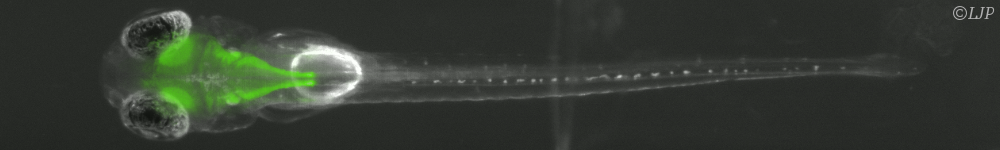
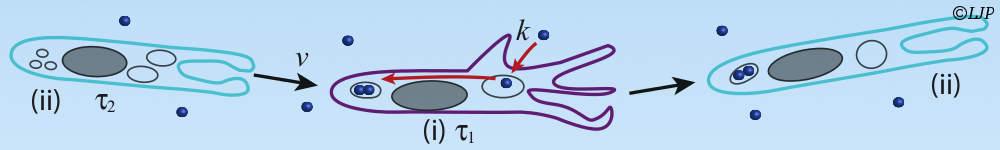
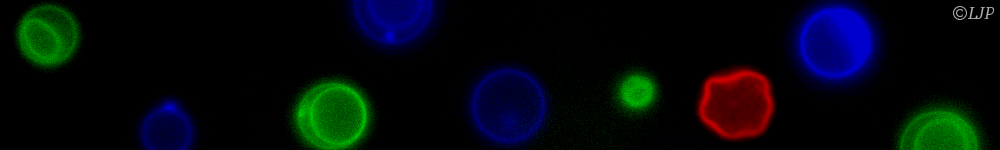
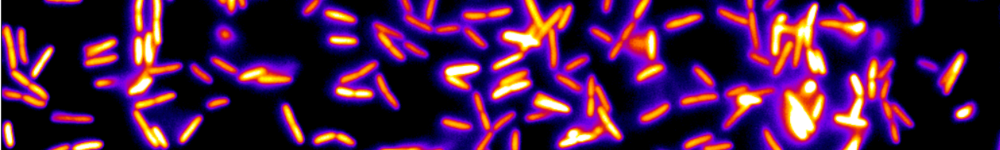
Microbes which live in hosts grow from small numbers to very large population sizes, and interact physically with their environment. Here is a summary of my biophysical modeling work, which uses a mix of analytical and numerical methods, and for some projects is in strong collaboration with experimentalists. This summary has two main parts. The first is focused on within-host dynamics, particularly about the bacterial dynamics within the digestive tract. With stochastic models of population dynamics, we developed methods to infer biologically interesting parameters of the population dynamics from the distribution of tagged populations. One of the results we contributed to is that one way for antibodies to protect the host against pathogenic bacteria is by enchaining together bacterial daughter cells upon replication. We then showed that the time scale of replication and the time scale of the breaking of the antibody mediated links between bacteria, may interact such that only fast replicating bacteria, the most likely to be destabilizing the normal microbiota, get trapped in antibody-mediated clusters. The second part of this summary is focused on minimal models exploring the impact of population structure on microbial evolution. We investigated the effect of antibody-mediated binding, reducing genetic diversity at transmission, on bacterial adaptation, using a multi-scale model, integrating the effects of growth within the host, and transmission between hosts. Another aspect is that in the digestive tract, the hydrodynamic flow may structure spatially the bacterial population, and we found that it results in a higher fixation probability for neutral mutants compared to the well-mixed case. This summary is concluded by a short overview of my near future research directions.